Cluster analysis of gene expression
In order to discover gene behaviors at the phenotype level by connecting genotype to phenotype, additional analyses are required. Cluster analysis, an unsupervised learning method, is a popular method that searches for patterns of gene expression associated with an experimental condition. Numerous clustering algorithms have been developed to address the problem of cluster analysis of gene expression data.
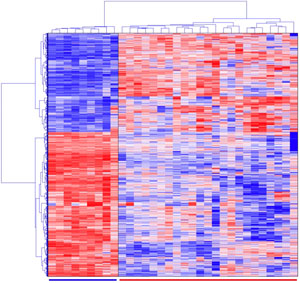
Hierarchical clustering results in a tree structure, where genes on the same branch at the desired level are considered to be in the same cluster. While this structure provides a rich visualization of the gene clusters, it has the limitation that a gene can be assigned only to one branch, i.e., one cluster. This is not always biologically reasonable because most, if not all, genes have multiple functions. In addition, because of the mechanism of one-way assignment of genes to branches, the results may not be globally optimal.
The alternative approach, partitioning clustering, includes two major methods, heuristic-based and model-based. The former assigns objects to clusters using a heuristic mechanism, while the latter uses quantifying uncertainty measures. Probability and possibility are the two uncertainty measures commonly used. While probabilistic bodies of evidence consist of singletons, possibilistic bodies of evidence are families of nested sets. Both probability and possibility measures are uniquely represented by distribution functions, but their normalization requirements are very different. Values of each probability distribution are required to add to 1, while for possibility distributions, the largest values are required to be 1. Moreover, the latter requirement may even be abandoned when possibility theory is formulated in terms of fuzzy sets. The mixture model with the expectation-maximization (EM) algorithm is a well-known method using the probability-based approach. This method has the advantages of a strong statistical basis and a statistics-based model selection. However, the EM algorithm converges slowly, particularly at regions where clusters overlap and requires the data distribution to follow some specific distribution model. Because gene expression data are likely to contain overlapping clusters and do not always follow standard data distributions, (e.g., Gaussian, Chi-squared), the mixture model with the EM method is not appropriate. Fuzzy clustering using the most popular algorithm, Fuzzy C-Means (FCM), is another model-based clustering approach that uses the possibility measure. FCM both converges rapidly and allows assignment of objects to overlapping clusters using the fuzzifier factor, m, where 1≤m<∞. When the value of m changes, the cluster overlaps will change accordingly and there will be no overlapping regions in the clustering results when the value of m is equal to 1. However, similar to the EM-based method and the other partitioning methods, FCM has the problem of determining the correct number of clusters. Even if the cluster number is known a priori, FCM may provide different cluster partitions. Cluster validation methods are required to determine the optimal cluster solution. Unfortunately, the clustering model of FCM is a possibility-based one. There is no straightforward statistics-based approach to evaluate a clustering model except the cluster validity indices based on the compactness and separation factors of the fuzzy partitions. However, there are problems with these validity indices; both with scaling the two factors and with over-fit estimates. We have combined the advantages of the EM and FCM methods, where FCM plays the key role in clustering the data, and proposed a method to convert the clustering possibility model into a probability one and then to use the Central Limit Theorem to compute the clustering statistics and determine the data distribution model that best describes the dataset. We applied the Bayesian method with log-likelihood ratio and Akaike Information Criterion measure using the estimated distribution model for a novel validation method for fuzzy clustering partitions (Thanh Le et al.). REFERENCES
- Dubois D. and Prade M. H. (1988) Possibility theory: an approach to computerized processing of uncertainty, Plenum Press.
- Dubois D. and Prade M. H. (1980) Fuzzy sets and systems: Theory and applications, Mathematics in Science and Engineering, Accademic Press.
- Klir G.J. and Yuan B. (1995) Fuzzy Sets and Fuzzy Logic: Theory and Applications, Prentice Hall.
- Xu L. and Jordan M.I. (1996) On convergence properties of the EM algorithm for Gaussian Mixtures. Neural Computation, Vol. 8, pp. 409-1431.
- Ma S. and Dai Y. (2011) Principal component analysis based methods in bioinformatics studies, Bioinformatics, Vol. 10, pp.1-9.
- Bezdek J.C. (1981) Pattern Recognition with Fuzzy Objective Function Algorithms, Plenum Press, NewYork.
- Dunn J.C. (1974) A fuzzy relative of the ISODATA process and its use in detecting compact well-separated clusters, J. Cybernet, Vol. 3, pp. 32-57.
- Thanh Le and Katheleen Gardiner (2011) A validation method for fuzzy clustering of gene expression data, Proc. of the 2011 Intl' Conf. on Bioinformatics & Computational Biology, Vol. I, pp. 23–29.